HADDOCK (High Ambiguity Driven protein-protein DOCKing) is an integrative platform developed at Utrecht University for the modelling of biomolecular complexes. HADDOCK distinguishes itself from other docking methods in the fact that it can use a large variety of experimental and bioinformatics data to drive the docking process. HADDOCK can deal with a large class of modelling problems including protein-protein, protein-nucleic acids, protein-peptide and protein-ligand complexes.
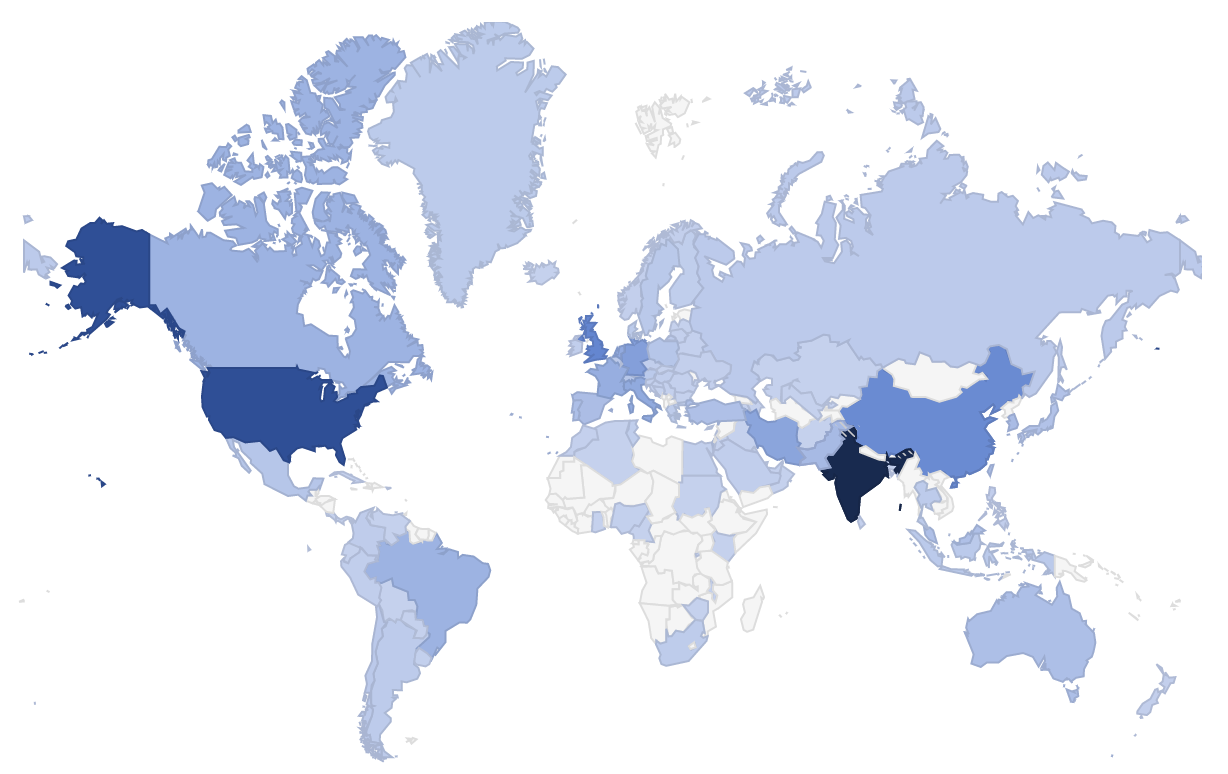
Since 2008 HADDOCK has been offered as a web portal, freely accessible to non-profit users (wenmr.science.uu.nl) as part of the WeNMR services. HADDOCK is also a thematic service in the European Open Science Cloud marketplace. This strategy has maximized the impact and valorisation of the research of our group (and the impact of NWO funding). Besides the application software, the service also provides automated pre- and post-processing, compute, temporary storage and job scheduling and monitoring for running the application, so that researchers do not need to worry about application porting and compute infrastructure. The portal is heavily used with >15500 registered users from >110 countries (see statistics), and receives a high number of citations.
We have obtained a NWO-ENW computing grant that allows us to maintain our services provisioning to serve the scientific community at large and maximise the impact of Dutch-funded research. Access to High Throughput Computing (HTC) grid resources is key. Not only does it allow to serve an international community, but it is also central to the research of the Utrecht group since significant computational recourses are required to keep developing new methods and model increasingly more complex and large macromolecular assemblies. The relevance of modelling biomolecular complexes is further highlighted by the fact that several major pharma companies are using our software. Our online services have also find their way into teaching as we see recurring registrations from students of various international universities.
SURFSara and the Dutch Dutch National Grid Initiative have always supported the computing required by the Utrecht portals, providing resources for more than 70% of the jobs that run last year. This computing grant allows us to ensure the continued operation of our portals by renewing our access to the Dutch HTC grid resources. For this, we have been granted 8 million SBU wall clock hours (4M per year).
