HADDOCK2.4 basic Antibody - Antigen tutorial including a comparison with AlphaFold
This tutorial consists of the following sections:
- Introduction
- Setup
- HADDOCK general concepts
- Inspecting the antibody and the identified paratope
- Inspecting the antigen stucture and NMR-identified binding site
- Using HADDOCK to model the antibody-antigen complex
- Analysing the results
- Visualisation
- BONUS 1: Does the antibody bind to a known interface of interleukin? ARCTIC-3D analysis
- BONUS 2: Modelling the antibody-antigen complex with AlphaFold2 - does it work?
- BONUS 3: Dissecting the interface energetics: PROT-ON analysis
- Conclusions
- Congratulations!
Introduction
This tutorial demonstrates the use of HADDOCK2.4 for predicting the structure of an antibody-antigen complex using NMR chemical shift perturbation data for the antigen and the knowledge of the hypervariable loops on the antibody as information to guide the docking.
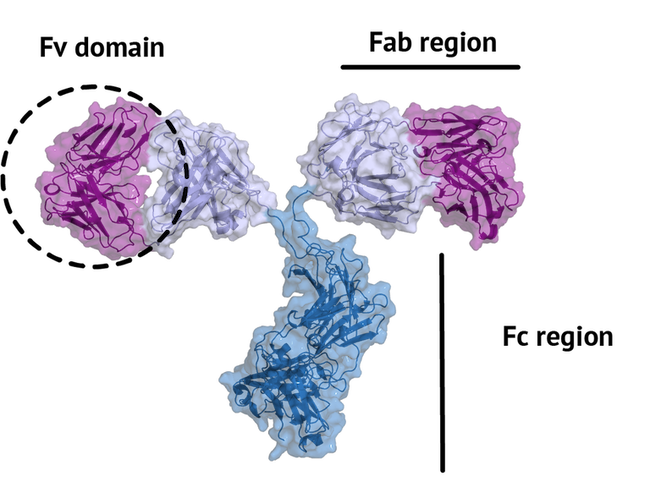
An antibody is a large protein that generally works by attaching itself to an antigen, which is a unique site of the pathogen. The binding harnesses the immune system to directly attack and destroy the pathogen. Antibodies can be highly specific while showing low immunogenicity, which is achieved by their unique structure. The fragment crystallizable region (Fc region) activates the immune response and is species-specific, i.e. the human Fc region should not induce an immune response in humans. The fragment antigen-binding region (Fab region) needs to be highly variable to be able to bind to antigens of various nature (high specificity). In this tutorial we will concentrate on the terminal variable domain (Fv) of the Fab region.
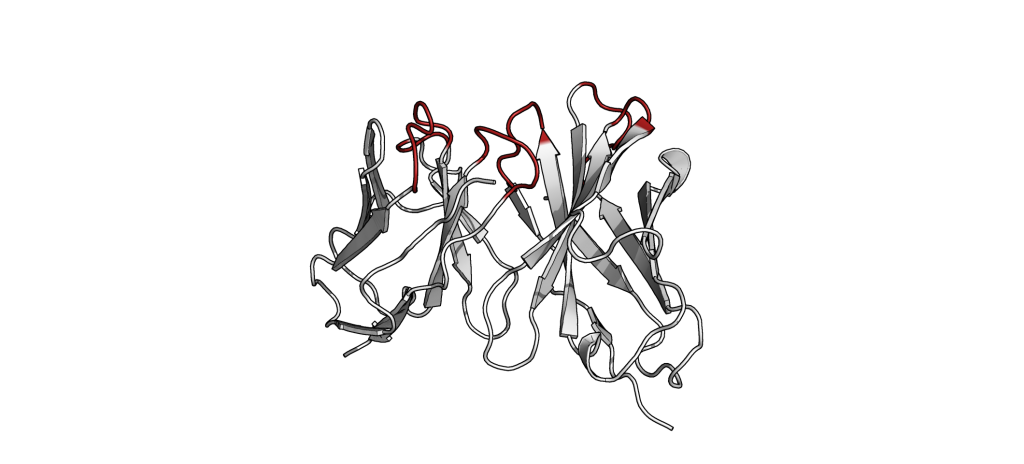
The small part of the Fab region that binds the antigen is called paratope. The part of the antigen that binds to an antibody is called epitope. The paratope consists of six highly flexible loops, known as complementarity-determining regions (CDRs) or hypervariable loops whose sequence and conformation are altered to bind to different antigens. CDRs are shown in red in the figure below:
In this tutorial we will be working with Interleukin-1β (IL-1β) (PDB code 4I1B)) as an antigen and its highly specific monoclonal antibody gevokizumab (PDB code 4G6K) (PDB code of the complex 4G6M).
We will make use of the HADDOCK2.4 webserver.
- R.V. Honorato, M.E. Trellet, B. Jiménez-García, J.J. Schaarschmidt, M. Giulini, V. Reys, P.I. Koukos, J.P.G.L.M. Rodrigues, E. Karaca, G.C.P. van Zundert, J. Roel-Touris, C.W. van Noort, Z. Jandová, A.S.J. Melquiond and A.M.J.J. Bonvin The HADDOCK2.4 web server: A leap forward in integrative modelling of biomolecular complexes. Nature Protoc, 19, 3219–3241 DOI: 10.1038/s41596-024-01011-0 (2024)
And our work on antibody antigen complexes is described in the following publications:
-
F. Ambrosetti, Z. Jandova and A.M.J.J. Bonvin. A protocol for information-driven antibody-antigen modelling with the HADDOCK2.4 webserver. Methods in molecular biology 2552, 267-282 (2023). Preprint.
-
F. Ambrosetti, T.H. Olsed, P.P. Olimpieri, B. Jiménez-García, E. Milanetti, P. Marcatilli and A.M.J.J. Bonvin. proABC-2: PRediction Of AntiBody Contacts v2 and its application to information-driven docking. Bioinformatics, 36, 5107–5108 (2020).
-
F. Ambrosetti, B. Jiménez-García, J. Roel-Touris and A.M.J.J. Bonvin. Modeling Antibody-Antigen Complexes by Information-Driven Docking. Structure, 28, 119-129 (2020). Preprint freely available from here.
Throughout the tutorial, coloured text will be used to refer to questions or instructions, and/or PyMOL commands.
This is a question prompt: try answering it! This an instruction prompt: follow it! This is a PyMOL prompt: write this in the PyMOL command line prompt!
Setup
In order to run this tutorial you will need to have the following software installed: PyMOL.
Also, if not provided with special workshop credentials to use the HADDOCK portal, make sure to register in order to be able to submit jobs. Use for this the following registration page: https://wenmr.science.uu.nl/auth/register/haddock.
Further we are providing pre-processed PDB files for docking and analysis. They have been processed (see below) to facilitate their use in HADDOCK and for allowing comparison with the known reference structure of the complex. For this download and unzip the following zip archive and note the location of the extracted PDB files in your system. You should find the following three files:
4G6K_fv.pdb: The PDB file of the unbound(free) form of the antibody with the two chains defined as a single chain and with residues renumbered to avoid overlap in numbering between the chains. The structure was further truncated to only keep the two domains involved in binding (to save computational time).4I1B-matched.pdb: The PDB file of the unbound(free) form of the antigen, renumbered to match the numbering of the reference complex.4G6M-matched.pdb: The PDB file of the reference antibody-antigen complex, matching the chainIDs and residue numbering of the free forms.
HADDOCK general concepts
HADDOCK (see https://www.bonvinlab.org/software/haddock2.4) is a collection of python scripts derived from ARIA (https://aria.pasteur.fr) that harness the power of CNS (Crystallography and NMR System – https://cns-online.org) for structure calculation of molecular complexes. What distinguishes HADDOCK from other docking software is its ability, inherited from CNS, to incorporate experimental data as restraints and use these to guide the docking process alongside traditional energetics and shape complementarity. Moreover, the intimate coupling with CNS endows HADDOCK with the ability to actually produce models of sufficient quality to be archived in the Protein Data Bank.
A central aspect to HADDOCK is the definition of Ambiguous Interaction Restraints or AIRs. These allow the translation of raw data such as NMR chemical shift perturbation or mutagenesis experiments into distance restraints that are incorporated in the energy function used in the calculations. AIRs are defined through a list of residues that fall under two categories: active and passive. Generally, active residues are those of central importance for the interaction, such as residues whose knockouts abolish the interaction or those where the chemical shift perturbation is higher. Throughout the simulation, these active residues are restrained to be part of the interface, if possible, otherwise incurring in a scoring penalty. Passive residues are those that contribute for the interaction, but are deemed of less importance. If such a residue does not belong in the interface there is no scoring penalty. Hence, a careful selection of which residues are active and which are passive is critical for the success of the docking. The docking protocol of HADDOCK was designed so that the molecules experience varying degrees of flexibility and different chemical environments, and it can be divided in three different stages, each with a defined goal and characteristics:
1. Randomization of orientations and rigid-body minimization (it0)
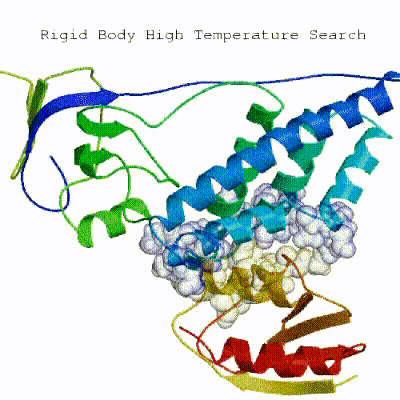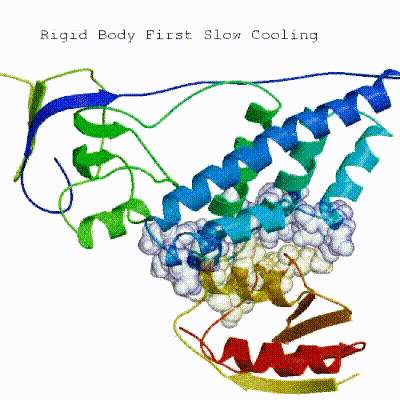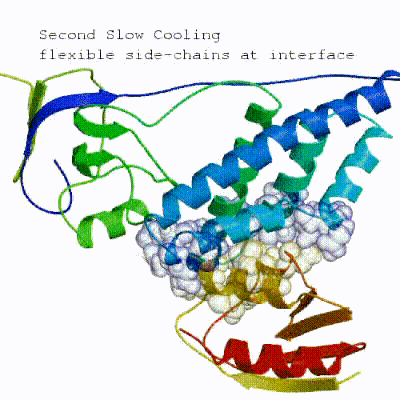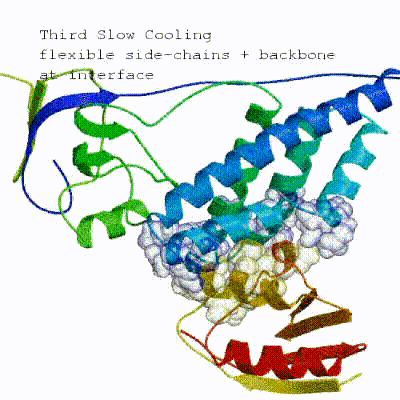
In this initial stage, the interacting partners are treated as rigid bodies, meaning that all geometrical parameters such as bonds lengths, bond angles, and dihedral angles are frozen. The partners are separated in space and rotated randomly about their centres of mass. This is followed by a rigid body energy minimization step, where the partners are allowed to rotate and translate to optimize the interaction. The role of AIRs in this stage is of particular importance. Since they are included in the energy function being minimized, the resulting complexes will be biased towards them. For example, defining a very strict set of AIRs leads to a very narrow sampling of the conformational space, meaning that the generated poses will be very similar. Conversely, very sparse restraints (e.g. the entire surface of a partner) will result in very different solutions, displaying greater variability in the region of binding.
See animation of rigid-body minimization (it0):

2. Semi-flexible simulated annealing in torsion angle space (it1)
The second stage of the docking protocol introduces flexibility to the interacting partners through a three-step molecular dynamics-based refinement in order to optimize interface packing. It is worth noting that flexibility in torsion angle space means that bond lengths and angles are still frozen. The interacting partners are first kept rigid and only their orientations are optimized. Flexibility is then introduced in the interface, which is automatically defined based on an analysis of intermolecular contacts within a 5Å cut-off. This allows different binding poses coming from it0 to have different flexible regions defined. Residues belonging to this interface region are then allowed to move their side-chains in a second refinement step. Finally, both backbone and side-chains of the flexible interface are granted freedom. The AIRs again play an important role at this stage since they might drive conformational changes.
See animation of semi-flexible simulated annealing (it1):
3. Refinement in Cartesian space with explicit solvent (water)
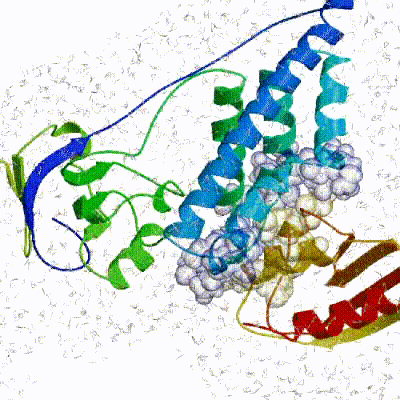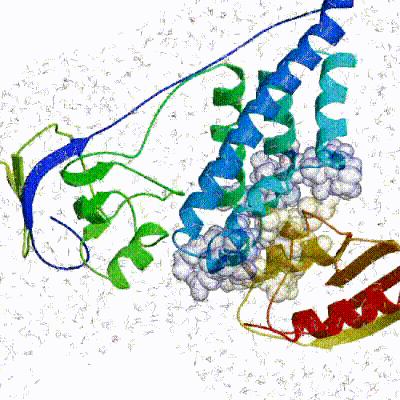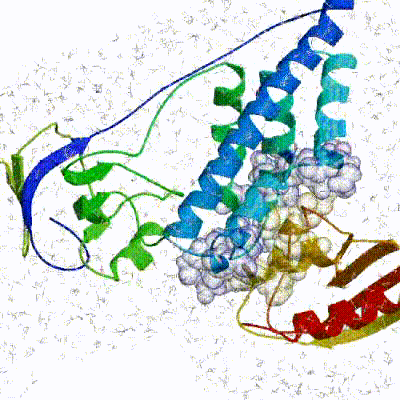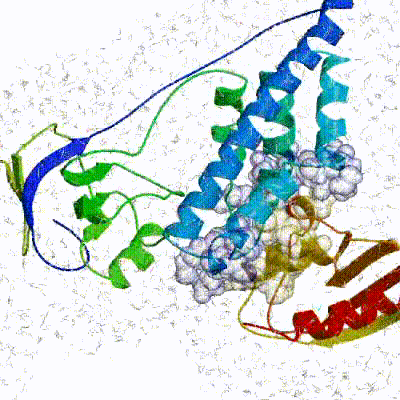
Note: This stage was part of the standard HADDOCK protocol up to (and including) v2.2. As of v2.4 it is no longer performed by default but the user still has the option of enabling it. In its place, a short energy minimisation is performed instead. The final stage of the docking protocol immerses the complex in a solvent shell so as to improve the energetics of the interaction. HADDOCK currently supports water (TIP3P model) and DMSO environments. The latter can be used as a membrane mimic. In this short explicit solvent refinement the models are subjected to a short molecular dynamics simulation at 300K, with position restraints on the non-interface heavy atoms. These restraints are later relaxed to allow all side chains to be optimized.
See animation of refinement in explicit solvent (water):
The performance of this protocol of course depends on the number of models generated at each step. Few models are less probable to capture the correct binding pose, while an exaggerated number will become computationally unreasonable. The standard HADDOCK protocol generates 1000 models in the rigid body minimization stage, and then refines the best 200 – regarding the energy function - in both it1 and water. Note, however, that while 1000 models are generated by default in it0, they are the result of five minimization trials and for each of these the 180º symmetrical solution is also sampled. Effectively, the 1000 models written to disk are thus the results of the sampling of 10.000 docking solutions.
The final models are automatically clustered based on a specific similarity measure - either the positional interface ligand RMSD (iL-RMSD) that captures conformational changes about the interface by fitting on the interface of the receptor (the first molecule) and calculating the RMSDs on the interface of the smaller partner, or the fraction of common contacts (FCC) (current default) that measures the similarity of the intermolecular contacts. For RMSD clustering, the interface used in the calculation is automatically defined based on an analysis of all contacts made in all models.
Inspecting the antibody and the identified paratope
Nowadays there are several computational tools that can identify the paratope (the residues of the hypervariable loops involved in the interaction) from the provided antibody sequence. In this tutorial we will use data obtained with the ProABC-2 server developed in our group. ProABC-2 uses a convolutional neural network to identify not only residues which are located in the paratope region but also the nature of interactions they are most likely involved in (hydrophobic or hydrophilic). The work is described in Ambrosetti, et al Bioinformatics, 2020. Details on how to run ProABC-2 for the antibody in this tutorial can be found here.
The list of of predicted paratope residues (matching the numbering of the HADDOCK-ready PDB file) is:
26,27,28,29,30,31,32,55,56,57,101,102,103,106,108,146,147,148,150,151,152,170,172,212,213,214,215
Note: Antibodies consist of two chains (the Heavy and Light chains, with corresponding chainID (H/L). They also have the peculiarity that some residues in the hypervariable loops are denotated as insertions, meaning that they have the same residue number as another residue, but with an additional A, B, … letter to the residue number to denote the insertion. For use in HADDOCK, these insertions must be removed, i.e. renumbered to have a sequential numbering of residues. Further, the two chains should be treated as a single chain with non-overlapping residue numbering. For this tutorial we are providing a HADDOCK-ready file where this has already been done. To see details on how to preprocess an antibody structure for use in HADDOCK refer to our more advanced antibody-antigen tutorial. In case of insertions, pdb_fixinsert from PDB-Tools can be used.
We will now inspect the antibody structure. For this start PyMOL and load the HADDOCK-ready PDB file of the antibody:
File menu -> Open -> select 4G6K_fv.pdb
We will now highlight the predicted paratope. In PyMOL type the following commands:
Let us now switch to a surface representation to inspect the predicted binding site.
Inspect the surface.
Do the identified residues form a well defined patch on the surface?
See surface view of the paratope:
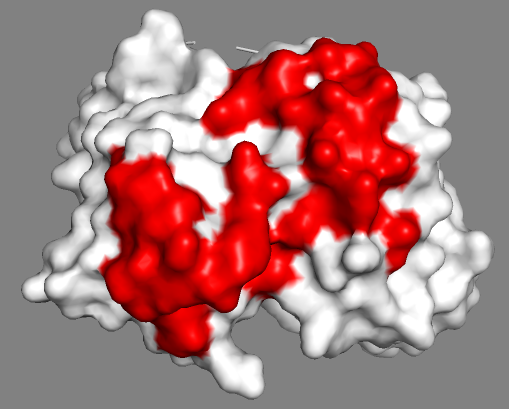

Inspecting the antigen stucture and NMR-identified binding site
The article describing the crystal structure of the antibody-antigen complex we are modelling also reports on experimental NMR chemical shift titration experiments to map the binding site of the antibody (gevokizumab) on Interleukin-1β. The residues affected by binding are listed in Table 5 of Blech et al. JMB 2013:
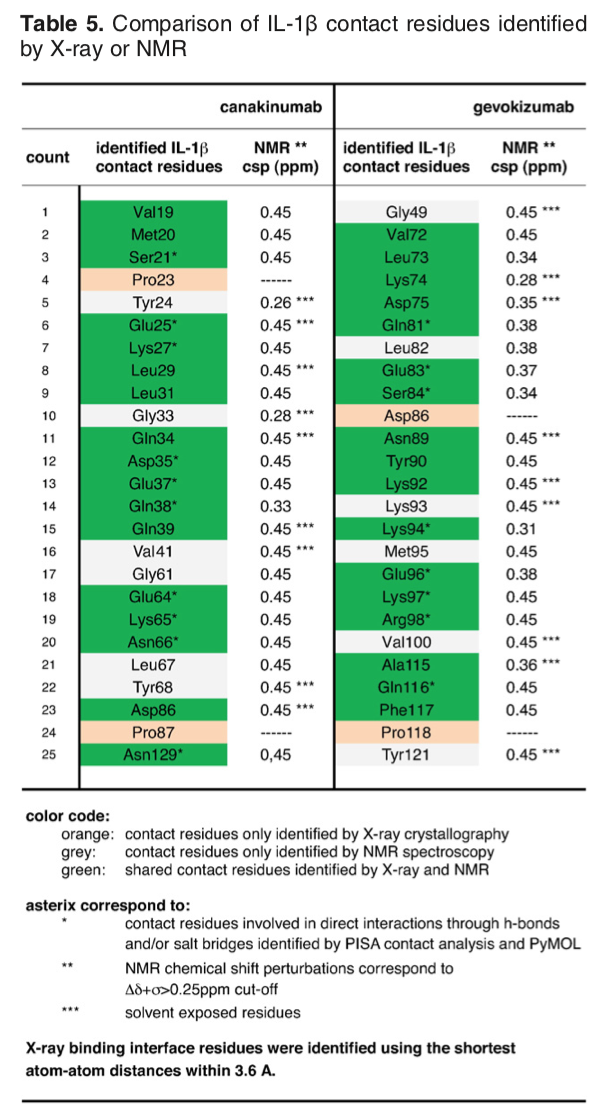
Note that the structure we are using for the docking has its residue numbering shifted by -2. The list of binding site (epitope) residues identified by NMR (corrected for the shift in numbering) is:
70,71,72,73,81,82,87,88,90,92,94,95,96,113,114,115
We will now visualize the epitope on Interleukin-1β. For this start PyMOL and from the PyMOL File menu open the provided PDB file of the antigen.
File menu -> Open -> select 4I1B-matched.pdb
Inspect the surface.
Do the identified residues form a well defined patch on the surface?
The answer to that question should be yes, but we can see some residues not colored that might also be involved in the binding (there are some white spots around/in the red surface.
See surface view of the epitope identified by NMR
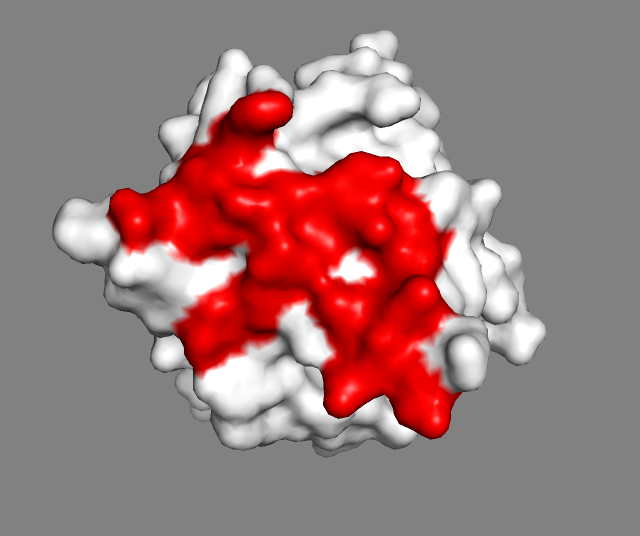
Note that in HADDOCK we are dealing with potentially uncomplete binding sites by defining surface neighbours as passive residues. These are added to the definition of the interface but will not lead to any energetic penalty if they are not part of the binding site in the final models, while the residues defined as active (typically the identified or predicted binding site residues) will. When using the HADDOCK server, passive residues will be automatically defined (default option - this can be turned off).
Using HADDOCK to model the antibody-antigen complex
Registration / Login
In previous steps we have identified the paratope and epitope residues of the antibody and antigen. Those can now be used to guide the docking. We will use this information to setup the docking.
If not provided with special workshop credentials, in order to start the submission you need first to register. For this go to https://wenmr.science.uu.nl/haddock2.4/ and click on Register.
To start the submission process, you are prompted for our login credentials. After successful validation of the credentials you can proceed to the structure upload under Submit a new job.
Note: The blue bars on the server can be folded/unfolded by clicking on the arrow on the left
Submission and validation of structures
We will make us of the HADDOCK 2.4 interface of the HADDOCK web server.
In this stage of the submission process we will upload the provided, pre-processed PDB structures.
-
Step 1: Define a name for your docking run in the field “Job name”, e.g. 4G6M-Ab-Ag-NMR.
-
Step 2: Select the number of molecules to dock, in this case the default 2.
-
Step 3: Input the first protein PDB file. For this, unfold the Molecule 1 - input if not already unfolded.
First molecule: where is the structure provided? -> “I am submitting it” Which chain to be used? -> All PDB structure to submit -> Browse and select 4G6K_fv.pdb
Note: Leave all other options to their default values.
- Step 4: Input the second protein PDB file. For this, unfold the Molecule 2 - input if not already unfolded.
First molecule: where is the structure provided? -> “I am submitting it” Which chain to be used? -> All PDB structure to submit -> Browse and select 4I1B-matched.pdb (the file you saved)
- Step 5: Click on the Next button at the bottom left of the interface. This will upload the structures to the HADDOCK webserver where they will be processed and validated (checked for formatting errors). The server makes use of Molprobity to check side-chain conformations, eventually swap them (e.g. for asparagines) and define the protonation state of histidine residues.
Definition of interfaces to guide the docking
If everything went well, the interface window should have updated itself and it should now show the list of residues for molecules 1 and 2. We will be making use of the text boxes below the residue sequence of every molecule to specify the list of active residues to be used for the docking run.
- Step 6: Specify the active residues for the first molecule. For this, unfold the “Molecule 1 - parameters” if not already unfolded.
Then uncheck the option to automatically define passive residues as for the antibody hypervariable loops this is not required.
Automatically define passive residues around the active residues -> uncheck (checked by default)
Note that HADDOCK with its default settings will automatically filter our residues that have a relative solvent surface accessibility below 15%.
Note: The web interface allows you to visualize the selected active residues.
- Step 7: Specify the residues for the second molecule. For this, unfold the “Molecule 2 - parameters” if not already unfolded.
Active residues -> 70,71,72,73,81,82,87,88,90,92,94,95,96,113,114,115
- Step 8: Click on the Next button on the bottom of the page. Since we have defined interface on both interaction partners, we can keep the default sampling parameters.
Job submission
This interface allows us to modify many parameters that control the behaviour of HADDOCK, but in our case, the default values are all appropriate. It also allows us to download the input structures of the docking run (in the form of a tgz archive) and a job_params file which contains all the settings and input structures for our run (in json format). We strongly recommend downloading this file as it will allow you to repeat the run by uploading it into the file upload interface of the HADDOCK webserver. The job_params file also serves as a run input reference. It can be edited to change a few parameters and repeat the run without going through the whole menu process again. An excerpt of this file is shown here:
It can serve as input reference for the run. This file can also be edited to change a few parameters for example. An excerpt of this file is shown here:
{
"amb_cool1": 10.0,
"amb_cool2": 50.0,
"amb_cool3": 50.0,
"amb_firstit": 0,
"amb_hot": 10.0,
"amb_lastit": 2,
"anastruc_1": 200,
...
}
- Step 9: Click on the “Submit” button at the bottom left of the interface.
Upon submission you will be presented with a web page which also contains a link to the previously mentioned haddockparameter file as well as some information about the status of the run.
Currently your run should be queued but eventually its status will change to “Running”:
The page will automatically refresh and the results will appear upon completion of the run (which can take between 1/2 hour to several hours depending on the size of your system and the load of the server). You will be notified by email once your job has successfully completed.
Analysing the results
Once your run has completed you will be presented with a result page showing the cluster statistics and some graphical representation of the data (and if registered, you will also be notified by email). When special course credentials, the number of models generated will have been automatically decreased (250/50/50) to allow the runs to complete within a reasonable amount of time.
In case you do not want to wait for your runs to be finished, a precalculated run can be found here.
How many clusters are generated?
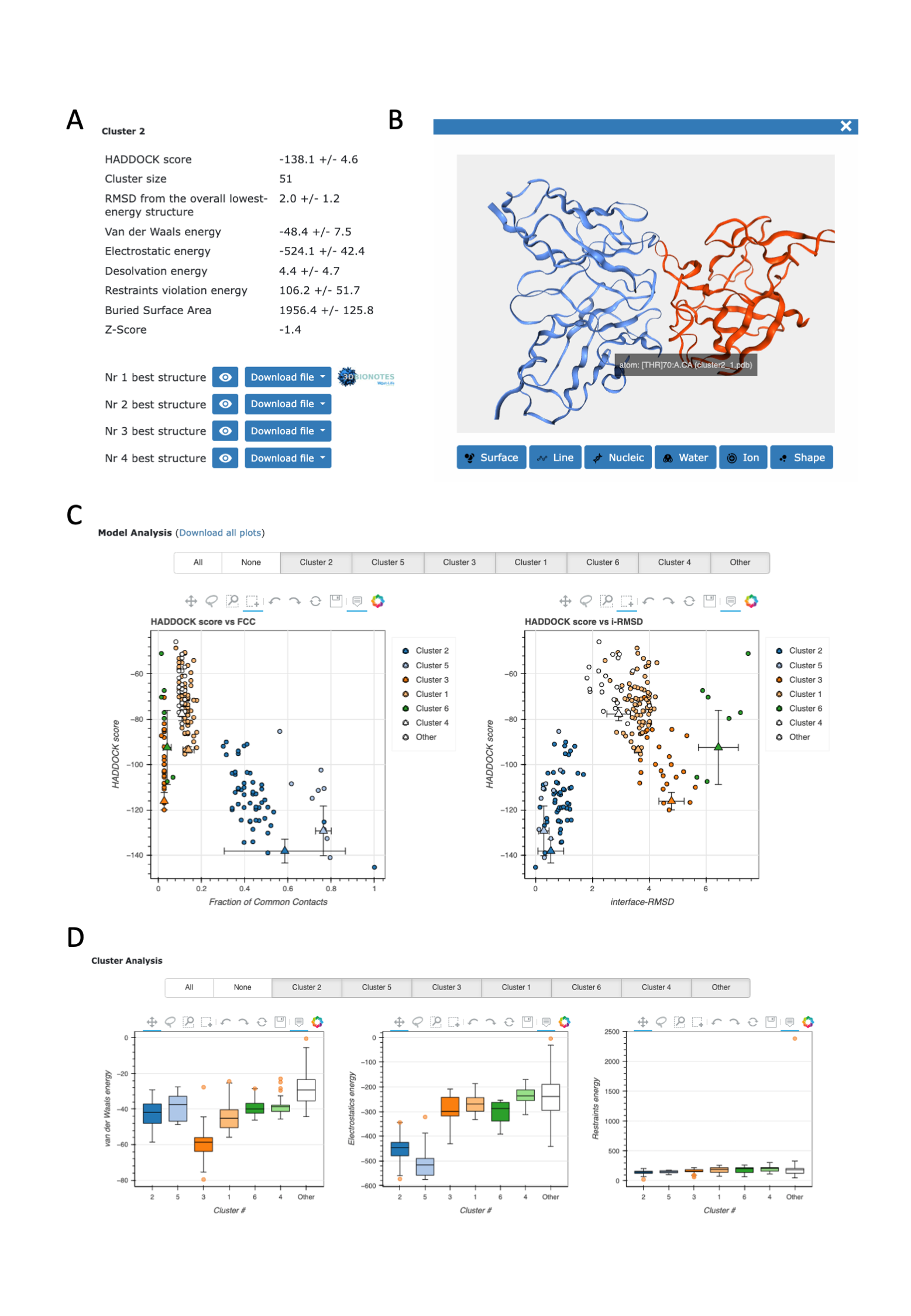
In the figure below you can see different parts of the result page.
In A the result page reports the number of clusters and for the top 10 clusters also the related statistics (HADDOCK score, Size, RMSD, Energies, BSA and Z-score). While the name of the clusters is defined by their size (cluster 1 is the largest, followed by cluster 2 etc..) the top 10 clusters are selected and sorted according to the average HADDOCK score of the best 4 models of each cluster, from the lowest (best) HADDOCK score to the highest (worst).
In B the visualization option of the various models is shown. You can visualize online a model by clicking on the eye icon, or download those for further analysis.
In C, model analysis a view of some graphical representation of the results shown at the bottom of the page under Model analysis is shown. Distribution of various measures (HADDOCK score, van der Waals energy, …) as a function of the Fraction of Common Contact with- and RMSD from the best generated model (the best scoring model) are shown. The models are color-coded by the cluster they belong to. You can turn on and off specific clusters, but also zoom in on specific areas of the plot.
In D, Cluster_analysis the distribution of components of the HADDOCK score (Evdw, Eelec and Edesol) for the various clusters is visualized.
The ranking of the clusters is based on the average score of the top 4 members of each cluster. The score is calculated as:
HADDOCKscore = 1.0 * Evdw + 0.2 * Eelec + 1.0 * Edesol + 0.1 * Eair
where Evdw is the intermolecular van der Waals energy, Eelec the intermolecular electrostatic energy, Edesol represents an empirical desolvation energy term adapted from Fernandez-Recio et al. J. Mol. Biol. 2004, and Eair the AIR energy.
Consider the cluster scores and their standard deviations. Is the top ranked cluster significantly better than the second one? (This is also reflected in the z-score).
In case the scores of various clusters are within standard deviation from each other, all should be considered as a valid solution for the docking. Ideally, some additional independent experimental information should be available to decide on the best solution.
Note: The type of calculations performed by HADDOCK does have some chaotic nature, meaning that you will only get exactly the same results unless you are running on the same hardware, operating system and using the same executable. The HADDOCK server makes use of EGI/EOSC high throughput computing (HTC) resources to distribute the jobs over a wide grid of computers worldwide. As such, your results might look slightly different from what is presented in the example output pages. That run was run on our local cluster. Small differences in scores are to be expected, but the overall picture should be consistent.
Visualisation
In the CAPRI (Critical Prediction of Interactions) Méndez et al. 2003
experiment, one of the parameters used is the Ligand root-mean-square deviation (l-RMSD) which is calculated by superimposing
the structures onto the backbone atoms of the receptor (the antibody in this case) and calculating the RMSD on the backbone
residues of the ligand (the antigen). To calculate the l-RMSD it is possible to either use the software
Profit or PyMOL.
For the sake of convenience we have provided you with a renumbered reference structure 4G6M-matched.pdb (in the zip archive you downloaded (see Setup)).
Then start PyMOL and load each cluster representative, e.g.:
File menu -> Open -> select cluster5_1.pdb
Repeat this for the best model (clusterX_1.pdb) of each cluster and 4G6M-matched.pdb. Once all files have been loaded, type in the PyMOL command window:
show cartoon
util.cbc
hide lines
color yellow, 4G6M-matched
Let us then superimpose all models on chain A (receptor) of the first model and calculate RMSD of chain B (ligand):
Note: On some machines the pymol rms_cur command can fail due to a bug in the PyMOL software. In this case you can use the following command instead:
align cluster1_1, 1GGR, cycles=0
This will align the two structures based on the all-atom RMSD, different from the ligand-RMSD (l-RMSD) that you can calculate with rms_cur and the above commands.
See the L-RMSDs for clusters in both scenarios:
* 4G6M-Ab-Ag cluster5_1 HADDOCKscore [a.u.] = -132.4 +/- 12.9 ligand-RMSD = 7.26Å * 4G6M-Ab-Ag cluster2_1 HADDOCKscore [a.u.] = -131.5 +/- 1.7 ligand-RMSD = 2.19Å * 4G6M-Ab-Ag cluster3_1 HADDOCKscore [a.u.] = -110.4 +/- 3.0 ligand-RMSD = 24.69Å * 4G6M-Ab-Ag cluster1_1 HADDOCKscore [a.u.] = -91.1 +/- 1.2 ligand-RMSD = 11.38Å * 4G6M-Ab-Ag cluster4_1 HADDOCKscore [a.u.] = -68.5 +/- 1.2 ligand-RMSD = 17.72Å
In CAPRI, the L-RMSD value defines the quality of a model:
- incorrect model: L-RMSD>10Å
- acceptable model: L-RMSD<10Å
- medium quality model: L-RMSD<5Å
- high quality model: L-RMSD<1Å
What is the quality of these models? Did any model pass the acceptable threshold?
Are there more clusters of acceptable or better quality?
Let’s now check if the active residues which we have defined (the paratope and epitope) are actually part of the interface. In the PyMOL command window type:
Note: You can turn on and off a cluster by clicking on its name in the right panel of the PyMOL window.
Note: By default HADDOCK will randomly discard 50% of the restraints (defined per active residue) for each model generated as a way of dealing with false positive (wrongly identified interface residues). As such not all of the defined paratope/epitope residue need to be at the interface in the resulting models.
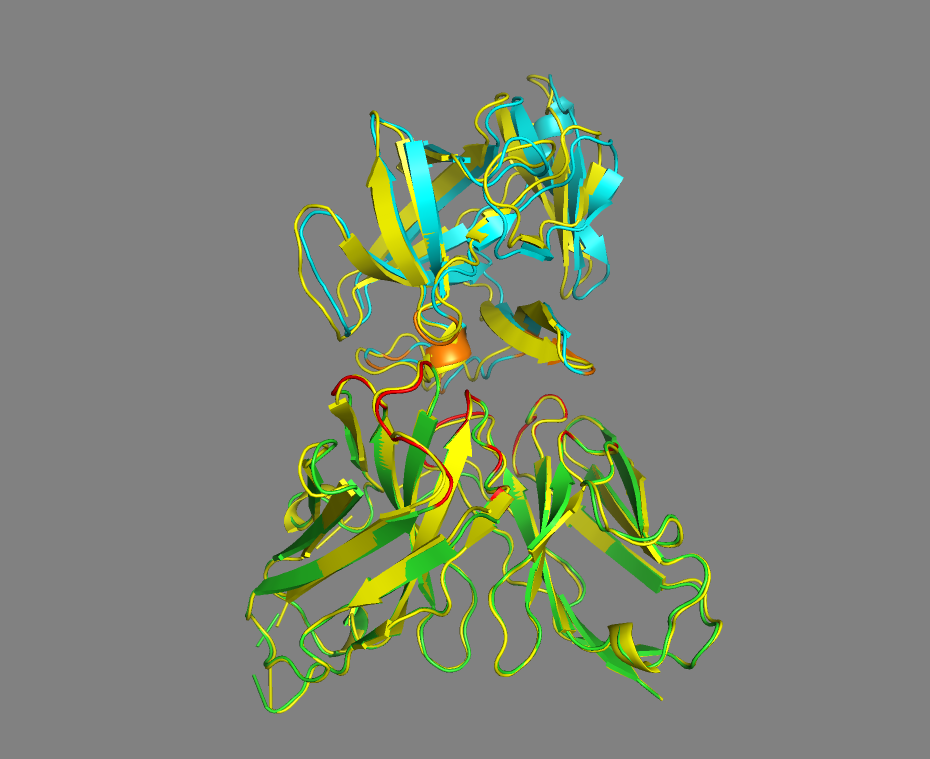
See the overlay of the best model onto the reference structure
Best HADDOCK model (cluster2) superimposed onto the reference crystal structure (in yellow)
BONUS 1: Does the antibody bind to a known interface of interleukin? ARCTIC-3D analysis
Gevokizumab is a highly specific antibody that targets an allosteric site of Interleukin-1β (IL-1β) in PDB file 4G6M, thus reducing its binding affinity for its substrate, interleukin-1 receptor type I (IL-1RI). Canakinumab, another antibody binding to IL-1β, has a different mode of action, as it competes directly with IL-1RI’s binding site (in PDB file 4G6J). For more details please check this article.
We will now use our new software, ARCTIC-3D, to visualize the binding interfaces formed by IL-1β. First, the program retrieves all the existing binding surfaces formed by IL-1β from the PDBe website. Then, these binding surfaces are compared and clustered together if they span a similar region of the selected protein (IL-1β in our case).
We now use the ARCTIC-3D web-server available here to run an ARCTIC-3D job targeting the uniprot ID of human Interleukin-1 beta, namely P01584.
Insert the selected uniprot ID in the UniprotID field.
Leave the other parameters as they are and click on Submit.
Wait a few seconds for the job to complete or access a precalculated run here.
How many interface clusters were found for this protein?
Once you download the output archive, you can find the clustering information presented in the dendrogram:
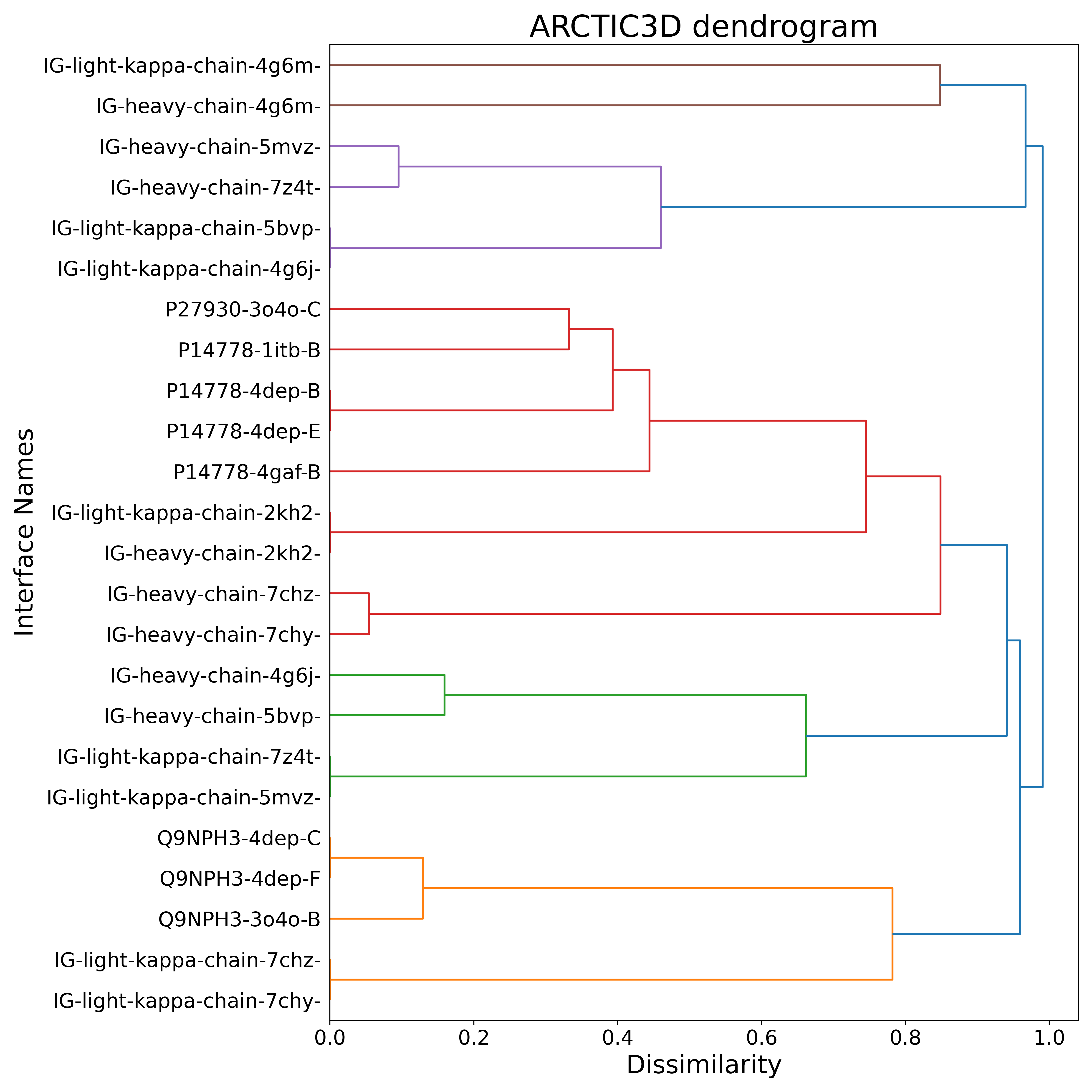
We can see how the two 4G6M antibody chains are recognized as a unique cluster, clearly separated from the other binding surfaces and, in particular, from those proper to IL-1RI (uniprot ID P14778).
Rerun ARCTIC-3D with a clustering threshold equal to 0.95
This means that the clustering will be looser, therefore lumping more dissimilar surfaces into the same object.
You can inspect a precalculated run here.
BONUS 2: Modelling the antibody-antigen complex with AlphaFold2 - does it work?
With the advent of Artificial Intelligence (AI) and AlphaFold we can also try to predict with AlphaFold this antibody-antigen complex. For a short introduction to AI and AlphaFold refer to this other tutorial introduction.
To predict our complex, we are going to use the AlphaFold2_mmseq2 Jupyter notebook which can be found with other interesting notebooks in Sergey Ovchinnikov’s ColabFold GitHub repository, making use of the Google Colab CLOUD resources.
Start the AlphaFold2 notebook on Colab by clicking here.
Note: The bottom part of the notebook contains instructions on how to use it.
Setting up the antibody-antigen complex prediction with AlphaFold2
To setup the prediction we need to provide the sequence of the heavy and light chains of the antibody and the sequence of the antigen. These are respectively
- Antibody heavy chain:
QVQLQESGPGLVKPSQTLSLTCSFSGFSLSTSGMGVGWIRQPSGKGLEWLAHIWWDGDESYNPSLKSRLTISKDTSKNQVSLKITSVTAADTAVYFCARNRYDPPWFVDWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKRVEP
- Antibody light chain:
DIQMTQSTSSLSASVGDRVTITCRASQDISNYLSWYQQKPGKAVKLLIYYTSKLHSGVPSRFSGSGSGTDYTLTISSLQQEDFATYFCLQGKMLPWTFGQGTKLEIKRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRG
- Antigen:
APVRSLNCTLRDSQQKSLVMSGPYELKALHLQGQDMEQQVVFSMSFVQGEESNDKIPVALGLKEKNLYLSCVLKDDKPTLQLESVDPKNYPKKKMEKRFVFNKIEINNKLEFESAQFPNWYISTSQAENMPVFLGGTKGGQDITDFTMQFVSS
To use AlphaFold2 to predict this antibody-antigen complex follow the following steps:
For your convenience the full sequence with colons is:
QVQLQESGPGLVKPSQTLSLTCSFSGFSLSTSGMGVGWIRQPSGKGLEWLAHIWWDGDESYNPSLKSRLTISKDTSKNQVSLKITSVTAADTAVYFCARNRYDPPWFVDWGQGTLVTVSS:DIQMTQSTSSLSASVGDRVTITCRASQDISNYLSWYQQKPGKAVKLLIYYTSKLHSGVPSRFSGSGSGTDYTLTISSLQQEDFATYFCLQGKMLPWTFGQGTKLEIK:VRSLNCTLRDSQQKSLVMSGPYELKALHLQGQDMEQQVVFSMSFVQGEESNDKIPVALGLKEKNLYLSCVLKDDKPTLQLESVDPKNYPKKKMEKRFVFNKIEINNKLEFESAQFPNWYISTSQAENMPVFLGGTKGGQDITDFTMQFVSS
Define the jobname, e.g. Ab_Ag
In the top section of the Colab, click: Runtime > Run All
(It may give a warning that this is not authored by Google, because it is pulling code from GitHub). This will automatically install, configure and run AlphaFold for you - leave this window open. After the prediction complete you will be asked to download a zip-archive with the results (if you configured it to use Google Drive, a result archive will be automatically saved to your Google Drive).
Time to grap a cup of tea or a coffee!
And while waiting try to answer the following questions:
How do you interpret AlphaFold’s predictions? What are the predicted LDDT (pLDDT), PAE, iptm?
Tip: Try to find information about the prediction confidence at https://alphafold.ebi.ac.uk/faq. A nice summary can also be found here.
Pre-calculated AlphFold2 predictions are provided here. This archive contains the fives predicted models (the naming indicates the rank), figures (png) files (PAE, pLDDT, coverage) and json files containing the corresponding values (the last part of the json files report the ptm and iptm values).
Analysis of the generated AF2 models
While the notebook is running models will appear first under the Run Prediction section, colored both by chain and by pLDDT.
The best model will then be displayed under the Display 3D structure section. This is an interactive 3D viewer that allows you to rotate the molecule and zoom in or out.
Note that you can change the model displayed with the rank_num option. After changing it execute the cell by clicking on the run cell icon on the left of it.
How similar are the five models generated by AF2?
Consider the pLDDT of the various models (the higher the pLDDT the more reliable the model).
While the pLDDT score is an overall measure, you can also focus on the interface score reported in the iptm score (value between 0 and 1).
See the confidence statistis for the five generated models
Model1: pLDDT 90.5, ptmscore 0.734 and iptm 0.626
Model2: pLDDT 90.5, ptmscore 0.741 and iptm 0.655
Model3: pLDDT 90.4, ptmscore 0.723 and iptm 0.622
Model4: pLDDT 91.0, ptmscore 0.715 and iptm 0.623
Model5: pLDDT 90.0, ptmscore 0.699 and iptm 0.599
Based on the iptm scores, would you qualify those models as reliable?
Note that in this case the iptm score reports on all interfaces, i.e. both the interface between the two chains of the antibody, and the antibody-antigen interface
Another usefull way of looking at the model accuracy is to check the Predicted Alignmed Error plots (PAE) (also refered to as Domain position confidence). The PAE gives a distance error for every pair of residues: It gives AlphaFold’s estimate of position error at residue x when the predicted and true structures are aligned on residue y. Values range from 0 to 35 Angstroms. It is usually shown as a heatmap image with residue numbers running along vertical and horizontal axes and each pixel colored according to the PAE value for the corresponding pair of residues. If the relative position of two domains is confidently predicted then the PAE values will be low (less than 5A - dark blue) for pairs of residues with one residue in each domain. When analysing your complex, the diagonal block will indicate the PAE within each molecule/domain, while the off-diaganal blocks report on the accuracy of the domain-domain placement.
Our antibody-antigen complex consists of three interfaces:
- The interface between the heavy and light chains of the antibody
- The interface between the heavy chain of the antibody and the antigen
- The interface between the light chain of the antibody and the antigen
See the PAE plots for the five generated models
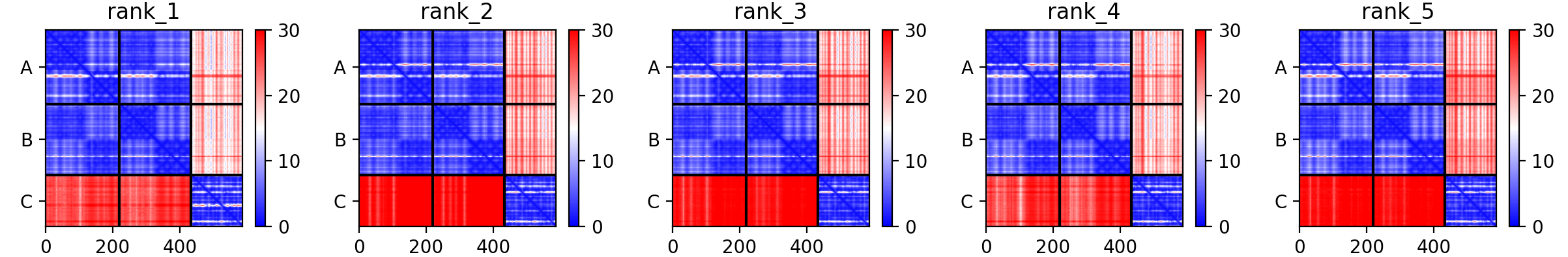
Based on the PAE plots, which interfaces can be considered reliable, unreliable?
Visualization of the generated AF2 models
Let’s now visualize the models in PyMOL. For this save your predictions to disk or download the precalculated AlphaFold2 model from here.
Start PyMOL and load via the File menu all five AF2 models (or use the command line to start PyMOL loading all models directly).
File menu -> Open -> select Ab_Ag_unrelaxed_rank_1_model_2.pdb
Repeat this for each model (Ab_Ag_unrelaxed_rank_X_model_Y.pdb or whatever the naming of your model is).
Let us superimpose all models on the antibody (the antibody in the provided AF2 models correspond to chains B and C):
util.cbc
select Ab_Ag_unrelaxed_rank_1_model_2 and chain B+C
alignto sele
This will align all clusters on the antibody, maximizing the differences in the orientation of the antigen.
Examine the various models. How does the orientation of the antigen differ between them?
Note: You can turn on and off a model by clicking on its name in the right panel of the PyMOL window.
See tips on how to visualize the prediction confidence in PyMOL
When looking at the structures generated by AlphaFold in PyMOL, the pLDDT is encoded as the B-factor.To color the model according to the pLDDT type in PyMOL:
spectrum b
Note that the scale in the B-factor field is the inverse of the color coding in the PAE plots: i.e. red mean reliable (high pLDDT) and blue unreliable (low pLDDT)). If you want to have the inverse coloring you can use a PyMol plugin, e.g. as described here.
Since we do have NMR chemical shift perturbation data for the antigen, let us check if the perturbed residues are at the interface in the AF2 models. Note that there is a shift in numbering of 2 residues between the AF2 and the HADDOCK models.
Does any model have the NMR-identified epitope at the interface with the antibody?
See the AlphaFold models with the NMR-mapped epitope
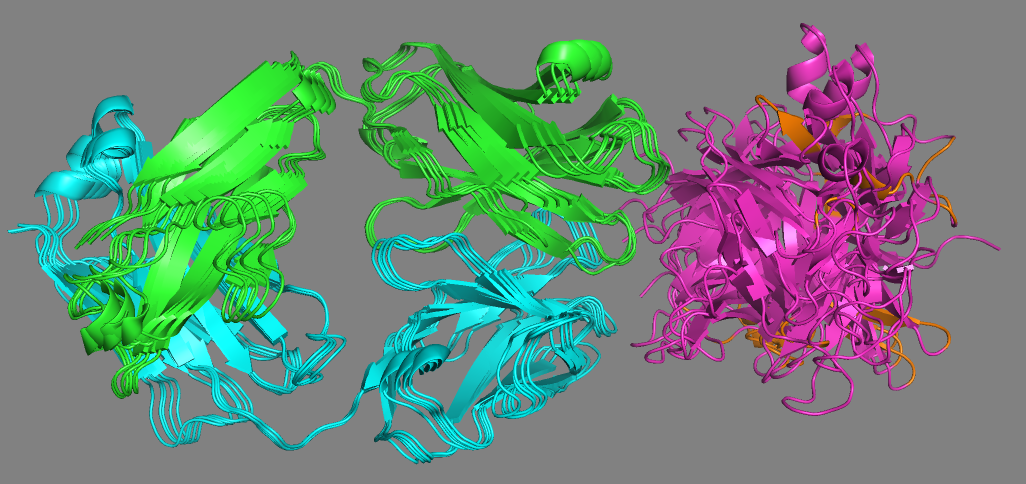
It should be clear from the visualization of the NMR-mapped epitope on the AF2 models that none does satisfy the NMR data. To further make that clear we can compare the models to the crystal structure of the complex (4G6M).
For this download and superimpose the models onto the crystal structure in PyMOL with the following commands:
fetch 4G6M
remove resn HOH
color yellow, 4G6M
select 4G6M and chain H+L
alignto sele
See the AlphaFold models superimposed onto the crystal structure of the complex (4G6M)
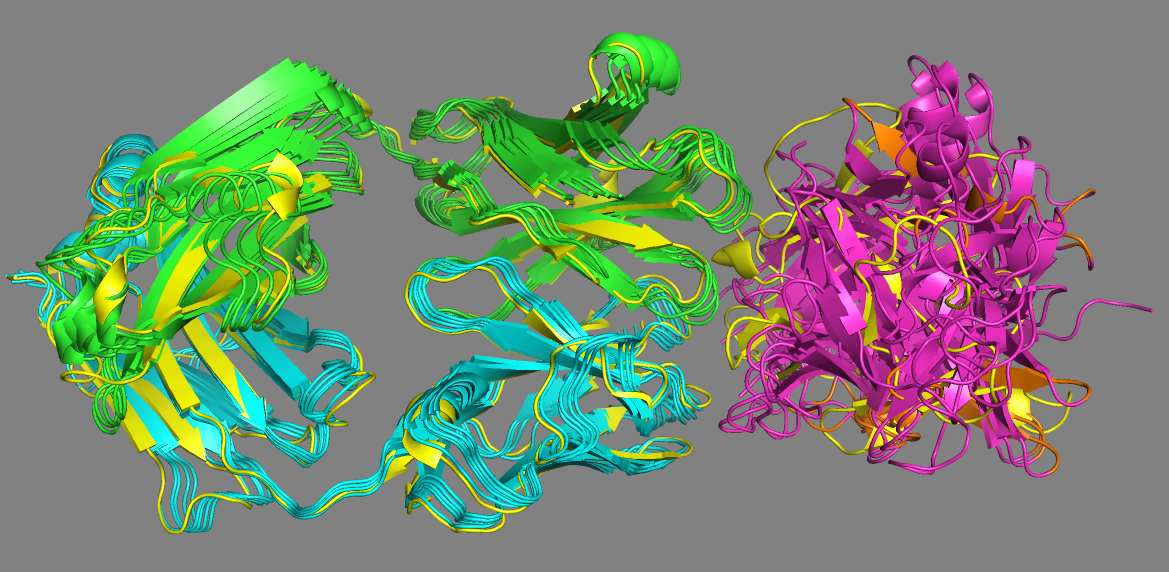
Finally, if you are curious, try running the same prediction using the new AlphaFold3 version. You can access the AlphaFold3 server here. You will however need a Google account to be able to use it.
BONUS 3: Dissecting the interface energetics: PROT-ON analysis
PROT-ON (Structure-based detection of designer mutations in PROTein-protein interface mutatiONs) is a tool and online server that allows to identify critical amino acids for redesigning a structurally characterized protein-protein interfaces, which paves the way for developing protein-based therapeutics to deal with diverse range of diseases. To find such amino acids positions, the residues across the protein interaction surfaces are either randomly or strategically mutated. Scanning mutations in this manner is experimentally costly. Therefore, computational methods have been developed to estimate the impact of an interfacial mutation on protein-protein interactions. PROT-ON scans all possible interfacial mutations by using EvoEF1 (active in both on the web server and stand-alone versions) or FoldX (active only in the stand-alone version) with the aim of finding the most mutable positions. The original publication describing PROT-ON can be found here.
Here we will use PROT-ON to analyse the interface of our antibody-antigen complex. We will use for that the provide matched reference structure (4G6M-matched) in which both chains of the antibody have the same chainID (A), which allows us to analyse all interface residues of the antibody at once.
Connect to the PROT-ON server page (link above)
Choose / Upload your protein complex –> select the provided 4G6M-matched.pdb file
Which dimer chains should be analyzed –> select chain A for the 1st molecule and B for the 2nd Pick the monomer for mutational scanning –> select the first molecule (the antibody) (switch the button)
You run should complete in 5 to 10 minutes. Once finished you will be presented with a result page summarizing the most deterious (lower the binding affinity) and most advantageous (increases the binding affinity) mutations.
Which possible mutations would you propose to improve the binding affinity of the antibody?
Inspect the proposed amino acids in PyMol. Can you rationalise why they might increase the affinity?
Now let’s consider how sensitive is this kind of analysis on the quality of the modelled complex. Instead of using the crystal structure, repeat this analysis using the best model of the top-ranked cluster (cluster5_1.pdb) or the best model of the second best cluster (overlapping scores), which has the lowest ligand-RMSD (cluster2_1.pdb).
Note: A pre-calculated run for the 4G6M crystal stucture is provided:
- 4G6M_matched (crystal structure)
Conclusions
We have demonstrated the modelling with HADDOCK of an antibody-antigen complex guided by knowledge of the paratope and epitope. Always check and compare multiple clusters, don’t blindly trust the cluster with the best HADDOCK score!
This example also illustrates that (for the time-being) AI-based prediction methods like AlphaFold2 still have issues with predicting this particular type of complexes. This should not come as a surprise as co-evolution (residues co-evolving at conserved locations) is an important source of information for AlphaFold2 and the antibody hypervariable loops are by definition not conserved and fastly mutating.
Congratulations!
You have completed this tutorial. If you have any questions or suggestions, feel free to contact us via email or asking a question through our support center.
And check also our education web page where you will find more tutorials!